Imaging Oxygen Dips in Mouse Cortex: Semi-NMF for Signal Extraction
We needed to detect dark, short-lived hypoxic regions in noisy mouse cortex videos, often spanning multiple consecutive frames, where simple thresholding failed. Semi-NMF gave us a robust way to isolate where these oxygen dips appear and how they evolve over time.
Key Takeaways
- Semi-NMF works well when preprocessing breaks strict non-negativity.
- Sparse spatial factors make transient “oxygen dips” easier to localize.
- Tuning
alphagives collaborators practical control over sensitivity.
Cover image from the related publication write-up.
Introduction
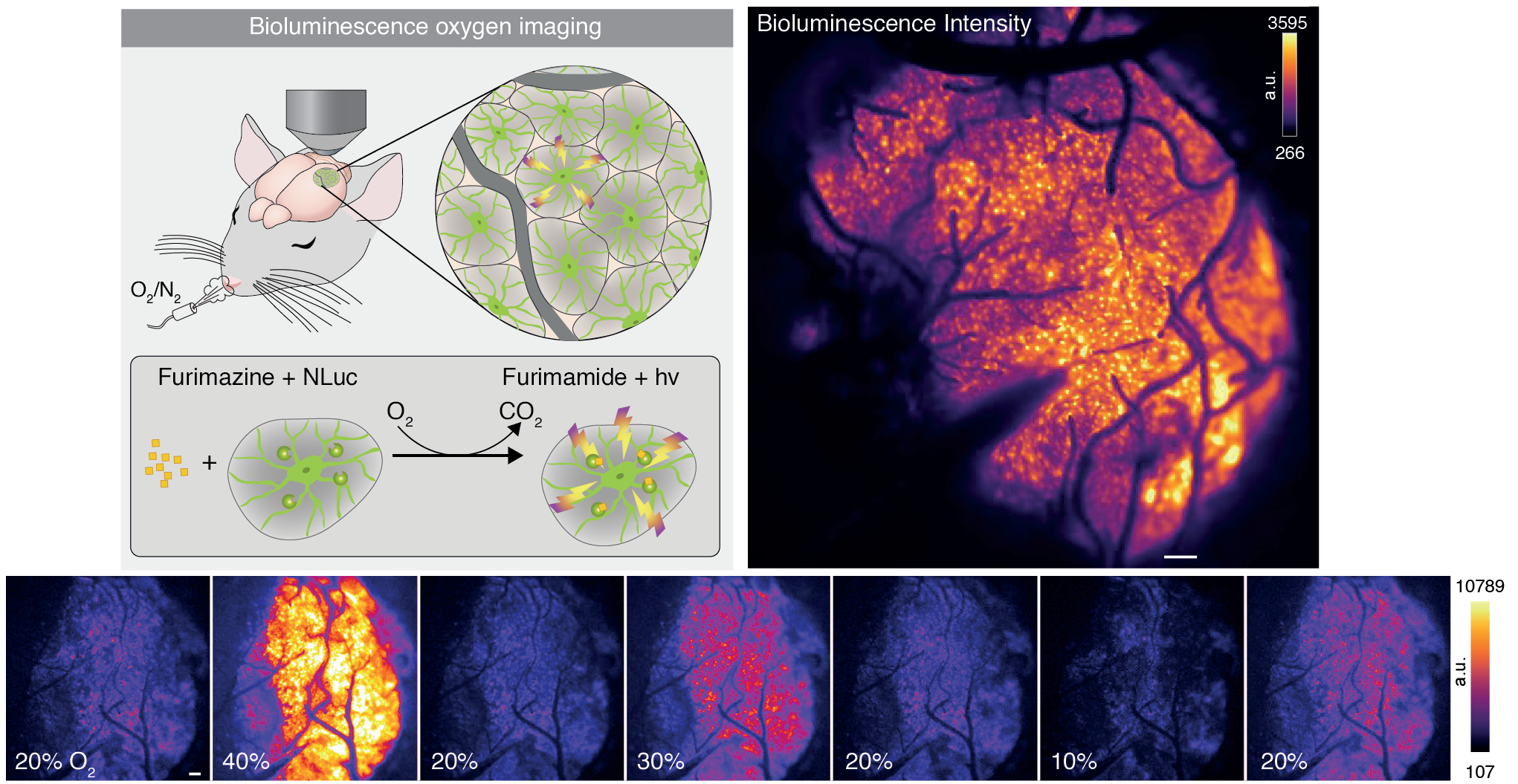
For my Bachelor’s thesis, I worked with the team at the Center for Translational Neuromedicine at the University of Copenhagen. The project revolved around imaging oxygen consumption in the mouse cortex. A specially designed molecule was injected into the brain; it emits light when it binds to oxygen. With a camera, we could then observe where and when oxygen was being used. Brighter pixels corresponded to more oxygen, darker ones to less. We started calling these darker, transient regions “oxygen dips,” while the team at Copenhagen referred to them as “hypoxic pockets” (they are the experts, after all!).
The Challenge
Detecting these pockets is difficult because bioluminescence imaging is inherently noisy. The “pockets” aren’t static; they are transient events that grow, migrate, and fade. Traditional thresholding fails here because the background intensity of the brain shifts over time.
Our approach
A video can be seen as a 3D matrix: two spatial dimensions and one temporal dimension, with each entry reflecting pixel intensity. This structure makes matrix factorization an appealing tool. After preprocessing steps like detrending and denoising, however, the data was no longer guaranteed to stay non-negative. That pushed us toward a semi-NMF setup.
We decomposed the recording into two components:
- a 2D spatial matrix (SP) marking where a dip appears, and
- a 1D temporal vector (ST) describing how the dip evolves in time.
We alternated between optimizing the spatial and temporal components until the reconstruction error stopped improving.
\[S_P = \arg\min_{S_P \in \mathbb{R}_+^{p}} \left\| S_{MT}^\top - S_T S_P^\top \right\|_F^2 + \alpha \|S_P\|_1\] \[S_T = \arg\min_{S_T \in \mathbb{R}^{p}} \left\| S_{MT} - S_P S_T^\top \right\|_F^2\]For the sparse spatial update, we used LassoLars, though other solvers would also have been suitable.
Result
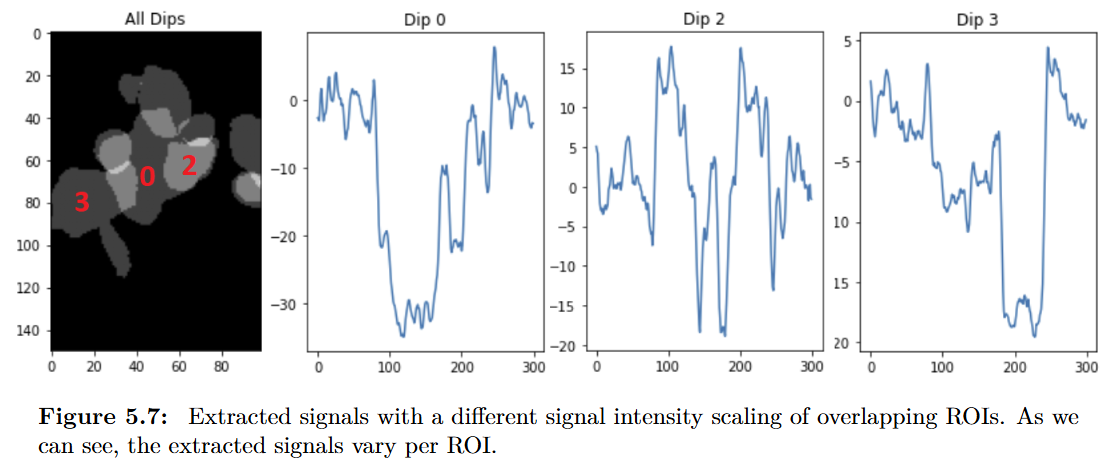
Signal Extraction and Ground Truth
The method reliably pulled out the transient dips-even when they overlapped or showed unusual shapes. The data was often noisy, and the dips themselves were not always well-defined, but the approach held up remarkably well. By adjusting the sparsity parameter $\alpha$, our collaborators could tune the sensitivity of the extraction: high values suppress weaker dips, while low values preserve more of them.
You can read our thesis here: Neuroscience Image Signal Segmentation and Extraction
Final Thoughts
Working with the Copenhagen team-and with our supervisor Prof. Dr. Michael Graber, who was consistently patient and supportive-was a highlight of this project. I’m proud that this work contributed to a publication in Science: Oxygen imaging of hypoxic pockets in the mouse cerebral cortex.
Dr. Felix Beinlich also published a detailed blog post about the work on his website: New Science Paper: Oxygen imaging of hypoxic pockets in the mouse cerebral cortex.
